| |
Top 10 FishMicrosat Families |
|
 |
|
 |
 |
 |
 |
| | |
| |
Top 10 FishMicrosat Species |
|
 |
|
 |
 |
 |
 |
|
|
|
Repeats statistics The program finds the frequency of different type of repeats from the records available in FishMicrosat and displays in the tabular and pie chart form.
Maximum repeats types AC 19793 | TG 16047 | CA 15018
| | Repeats | Frequency | Repeats | Frequency | TTTTA | 140 | TATTA | 128 | AAAAT | 126 | ATAAT | 116 | AAATA | 114 | ATTAT | 108 | TAAAA | 108 | ATATA | 98 | TAATA | 90 | TTTAT | 88 | AATAA | 82 | TATAT | 80 | TATTT | 76 | ATTTT | 76 | AAATT | 74 | TTATT | 74 | ATATT | 70 | TTATA | 70 | TATAA | 68 | ATAAA | 58 | AATAT | 58 | TATTC | 48 | TTCTA | 46 | AGAAT | 44 | TTTTG | 42 | TAGAA | 42 | AAAAC | 40 | ATTCT | 40 | GAATA | 36 | AATAG | 36 | TCTAT | 32 | TTATG | 32 | TTTAA | 30 | ATTAA | 30 | AATTA | 28 | TTTCT | 26 | CTATT | 24 | ATTAG | 24 | ATAGA | 22 | TTAAA | 22 | CAAAA | 20 | ATTTA | 20 | AAGAA | 20 | CTTTT | 20 | TTAGA | 18 | GAAGA | 18 | GTTTT | 18 | TTTGT | 18 | TTTAG | 18 | AAAAG | 16 | AGAGG | 16 | TCTCT | 16 | TTGAA | 16 | AATCT | 16 | AGAGA | 16 | ACAAA | 16 | AATTT | 16 | AAAGA | 16 | TTTTC | 16 | TAAAT | 14 | TTGTT | 14 | TAGTA | 14 | AAGTC | 14 | TGTTA | 14 | GAAAA | 14 | GAGAA | 14 | AAACA | 14 | ATAAC | 14 | CTCTT | 12 | AGAAA | 12 | TTAAT | 12 | TACTA | 12 | ATTCA | 12 | ATATG | 12 | AACAT | 12 | TATGT | 12 | TCTTC | 12 | CTCTC | 12 | AGTAT | 10 | TCTTT | 10 | TTGTA | 10 | ATGTT | 10 | AACTG | 10 | TTCTT | 10 | ATATC | 10 | AACAA | 10 | TTTCA | 10 | TTCTC | 10 | TCAAG | 10 | GAGGA | 10 | TAGTT | 8 | CATAT | 8 | ATTTC | 8 | ATCTA | 8 | AGATT | 8 | ACGAC | 8 | AGGTT | 8 | CCTTT | 8 | GACTT | 8 | TCTCC | 8 | GTTAT | 8 | AGAAG | 8 | TATAC | 8 | TCAAT | 8 | AAAGG | 8 | AACTA | 8 | TGACT | 8 | ATAGT | 8 | TAATT | 8 | GATAT | 8 | AACAC | 6 | TTATC | 6 | AAATG | 6 | CAAAG | 6 | TGTTC | 6 | GTTCA | 6 | TTCTG | 6 | CAAGT | 6 | TTTGC | 6 | GATGA | 6 | CAAAT | 6 | CTTCT | 6 | TGAAT | 6 | TCCTC | 6 | ACATA | 6 | AATTG | 6 | AAGAG | 6 | AAACT | 6 | AGGAA | 6 | ATGTA | 6 | GAGAG | 6 | GTGGT | 6 | ACTAA | 6 | ACAAC | 6 | TGTTT | 6 | AGTCA | 6 | TTGAC | 4 | ATGAA | 4 | GTGAA | 4 | GTTAC | 4 | TGAAG | 4 | AGATG | 4 | AGTTT | 4 | AATGT | 4 | TCATG | 4 | CTAAA | 4 | CTGTT | 4 | TAATC | 4 | TGTGT | 4 | ATTTG | 4 | ACGTG | 4 | TGATA | 4 | ACAGG | 4 | TCTCA | 4 | TAAAC | 4 | TGCTT | 4 | TATTG | 4 | GATGT | 4 | TATGA | 4 | GGGAA | 4 | AAGTG | 4 | AAATC | 4 | CTTGC | 4 | ATACT | 4 | ATCAC | 4 | CTTTA | 4 | TATAG | 4 | CTATA | 4 | GTTTA | 4 | GAACA | 4 | CTTTC | 4 | TTAGT | 4 | TATGG | 4 | CAGAC | 4 | GGAGA | 4 | CACTT | 4 | AATGA | 4 | CTCCT | 4 | TCCTT | 4 | TTACA | 4 | AGACA | 4 | TAAGA | 4 | GATTT | 4 | AGGAG | 4 | GAATG | 4 | AATCA | 4 | TCAAC | 4 | CCTCT | 4 | TAATG | 4 | AGACG | 4 | CATTG | 4 | AGGCA | 4 | TCAAA | 4 | ATGGA | 2 | TAGAG | 2 | CCAGT | 2 | ATGAC | 2 | TAACG | 2 | AAGCT | 2 | AATTC | 2 | GGCAG | 2 | TACTC | 2 | ATTGA | 2 | GAGAC | 2 | GCCGG | 2 | TCATT | 2 | ATTGT | 2 | GTAAA | 2 | GATGC | 2 | TTCAA | 2 | TAGGA | 2 | GACGT | 2 | TGCAA | 2 | GTCAA | 2 | CAGAA | 2 | CTCCA | 2 | AGATC | 2 | GGTTC | 2 | TGTCC | 2 | ATCCA | 2 | GATCA | 2 | TTTGA | 2 | TAGGC | 2 | GGACT | 2 | TGTTG | 2 | CTGTC | 2 | TTCAT | 2 | TAAGT | 2 | TTAGC | 2 | AGTGT | 2 | TGGTG | 2 | CAAGG | 2 | GCTCG | 2 | CACAA | 2 | TAGAT | 2 | GATTA | 2 | GTTCT | 2 | CATTA | 2 | AGAAC | 2 | AACAG | 2 | ACGGC | 2 | GATAA | 2 | CCCCT | 2 | TTCCT | 2 | TCGTA | 2 | ATGAT | 2 | GGGAC | 2 | ACAGA | 2 | TACTG | 2 | TTACC | 2 | AGACT | 2 | ACCAG | 2 | CGTCG | 2 | TACCC | 2 | TCTTG | 2 | ATGTC | 2 | AAGGC | 2 | TATCA | 2 | TCTGG | 2 | ATAGG | 2 | CAGGA | 2 | TGTGC | 2 | TAACA | 2 | TATCG | 2 | ATGCA | 2 | CTGAA | 2 | TCTGT | 2 | AAGAT | 2 | AATAC | 2 | AGCCG | 2 | GGAGG | 2 | CTATC | 2 | ATGGC | 2 | TTGAT | 2 | ATGCT | 2 | AACCT | 2 | TCGTG | 2 | TATGC | 2 | ATCAT | 2 | TTGCT | 2 | TTACT | 2 | CTCAC | 2 | AGTTG | 2 | AGGGA | 2 | CAAAC | 2 | GCGGG | 2 | GCCTT | 2 | GTGTA | 2 | GAATT | 2 | AACCA | 2 | AAGCA | 2 | CGTCT | 2 | TCTAA | 2 | ACCGA | 2 | CCTAT | 2 | TGCTG | 2 | GGGAG | 2 | GAGAT | 2 | TGATT | 2 | TTGCC | 2 | AAAGC | 2 | CCCCG | 2 | TTTCC | 2 | TACCT | 2 | ACGTA | 2 | ATTAC | 2 | AGCAG | 2 | GGCAA | 2 | GACAT | 2 | CTCTA | 2 | GTGTT | 2 | TCCAT | 2 | AAAGT | 2 | TGAGT | 2 | ACACA | 2 | CTCAA | 2 | GTGAC | 2 | GAAGG | 2 | GATCT | 2 | GAACC | 2 | CTAAC | 2 | AGTTC | 2 | CACAT | 2 | GCCTG | 2 | TACGT | 2 | CCTCA | 2 | GAGGC | 2 | CAATT | 2 | AAGGA | 2 | TGGAG | 2 | GTATC | 2 | ACACG | 2 | GATAC | 2 | TGTAT | 2 | AGGAC | 2 | GTATT | 2 | ACACT | 2 |
| |
| This pie chart repersents maximum frequencies of all types (mono, di, tri, tetera, penta and hexa) nucleotide repeats. 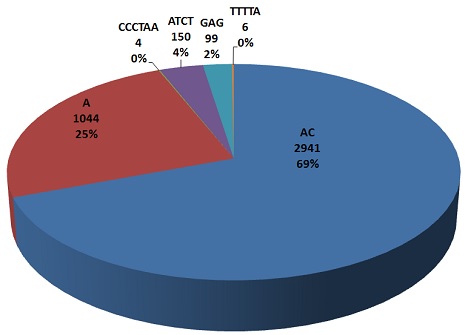
|
|
|